3-Dimensional Pharmaceuticals (3DP): Disruptive Drug Discovery Technology (1993-2003)
3-Dimensional Pharmaceuticals, Inc (3DP, Nasdaq DDDP) was a drug discovery company that I founded in 1993. 3DP was originally conceived with a focus on structure-based drug design, but almost immediately found itself in competition with a new generation of combichem and high throughput screening technology companies. To both establish credibility with the investment community and assist in financing operations, it was to imperative to form funded research collaborations with pharma partners, to be run in parallel with internal discovery programs. To achieve these goals, 3DP developed a comprehensive discovery platform that incorporated structure-based drug design, combichem directed through a comprehensive cheminformatics platform, and a unique, label-free, high throughput screening platform uniquely suited to screening modulators for targets identified using genomics methods. A discussion of these technologies, structure-based design, DirectedDiversity combinatorial Chemistry and Thermofluor screening technology are treated in the linked sections below.
What follows here are descriptions of the origins of 3DP as well as some of the events that impacted 3DP’s evolution as a biotechnology company until its eventual acquisition by John and Johnson in 2003.
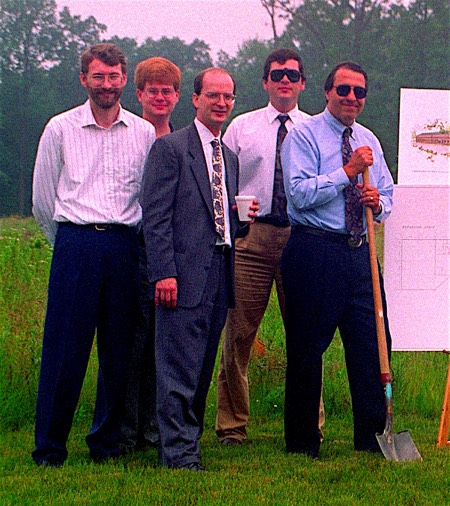
Roger Bone, Jim Myslik, Scott Horvitz, Rich Soll, and Ray Salemme at the Eagleview 3DP groundbreaking ceremony in 1994.
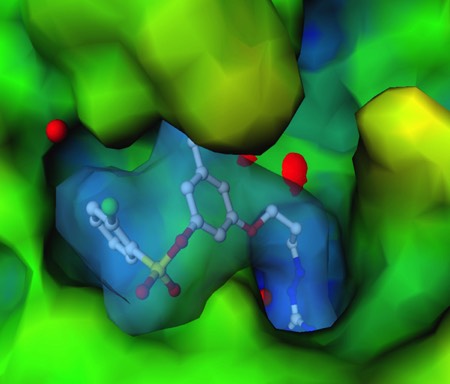
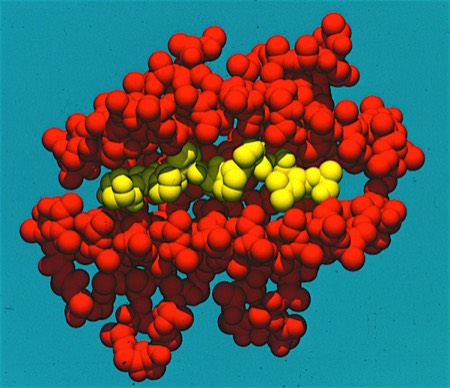
X-Ray Crystal Structure of 3DP Thrombin inhibitor bound in thrombin active site. 3DP used structure-based design strategy to generate a series of potent, orally active thrombin inhibitors. This program formed the basis for research and development collaborations with Wyeth Ayerst, and later Johnson & Johnson.
Work in the Wiley lab at Harvard had revealed the molecular features that Major Histocompatibility Complexes (MHCs) used to recognize antigenic peptides. The MHC peptide binding site seemed a suitable target for small-molecule structure-based design, although technical issues with MHC crystallography, and ultimately, discoveries relating to MHC biological function, prevented advancement of the program beyond its initial stages.
In 1995, 3DP was issued its first patent for “System and Method of Automatically Generating Chemical Compounds With Desired Properties”. The patent described a process that used computational methods to control of chemical synthesis of compounds with desired chemical properties, particularly using robotic chemical synthesis methods. At the time, this was a new concept that was very different from classical medicinal chemistry practice.
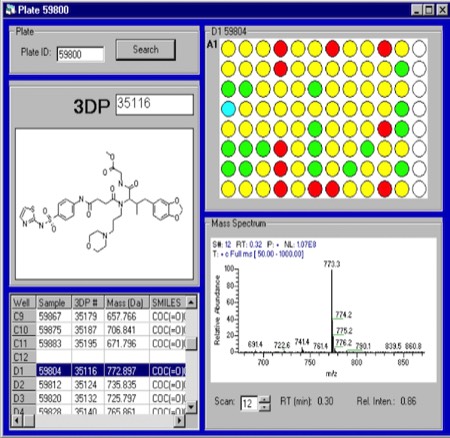
High-Throughput Compound Analysis and Purification at 3DP. 3DP synthesized several diverse compound libraries using a variety of synthetic chemistry platform approaches. It was quickly realized that post-screening analysis of reaction products that did not produce >90% of the desired yield product created a bottleneck in the hit definition process. Consequently, we developed a strategy where all synthesized compounds were submitted for HPLC-Mass spec analysis prior to screening, with all data tracked in the DirectedDiversity chemiformatics database. Compound QC and purification turned out to be a major investment area for 3DP, but one absolutely required to insure the fidelity of high-throughput discovery operations.
Cheminformatics for Combichem Compound Tracking. 3DP’s DirectedDiversity chemiformatics platform comprehensively tracked compound properties from synthesis through high-throughput screening and biological assay results.
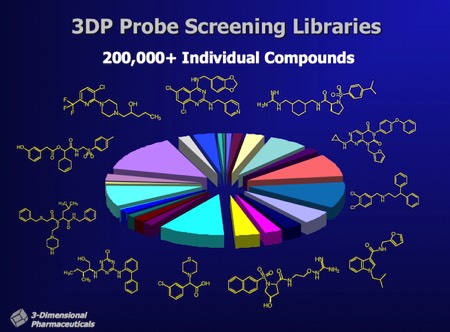
Compound Library Diversity at 3DP. 3DP’s experienced team of medicinal chemists collaborated with the DirectedDiversity computational team to generate novel, structurally diverse libraries containing hundreds of millions of compounds that were accessible through known chemical synthesis routes. Cheminformatics selections were then applied to drive synthesis of diverse representative subsets - typically containing several thousand compounds for incorporation in screening libraries. At any one time, 3DP maintained a screening library of ~50,000 to ~200,000 novel compounds for HTS screening.
Target Decryption using ThermoFluor. 3DP used cofactor and substrate “probe libraries” containing thousands of compounds to perform preliminary screens on novel drug discovery targets. Functional screens often revealed unknown dependencies on ions or cofactors, and in some cases could facilitate the functional identification of proteins with otherwise unknown biochemical function.
High Throughput Crystallization at 3DP. To support its structure-based drug design programs, 3DP established a high-throughput capability to screen crystallization conditions using robotic systems for both crystal tray setup and microscopic examination of crystal plate at periodic intervals. Crystallization trials were also facilitated using Thermofluor screens for crystallization conditions where the protein manifest optimal stability characteristics. The capability was an important asset that aided 3DP’s NIST ATP award to work in GPCR integral membrane protein structure.
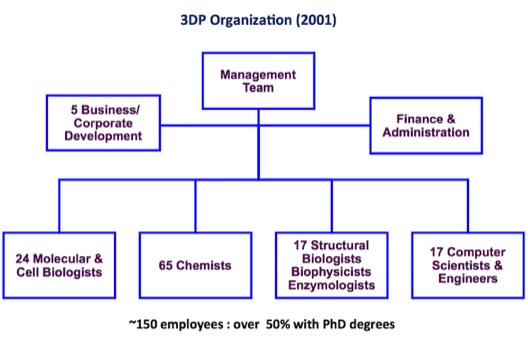
3DP Organization in 2001. 3DP was unusual for a biotechnology company owing to its relatively high complement of biophysicists, computational scientists and engineers. Although working as integrated teams at 3DP, capabilities were dispersed after 3DP’s acquisition by Johnson and Johnson in 2003.
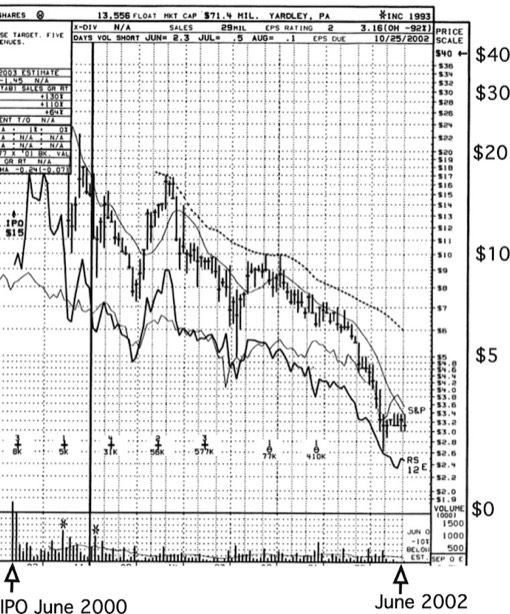
3DP went public on NASDAQ (DDDP) in a Bear Stearns-led IPO in 2000 at $15 a share, giving it an initial valuation of approximately $320 Million. Market enthusiasm transiently pushed the stock to a high of over $38 per share, before the stock began to decline owing to a combination of market forces, lack of top line revenue from collaborative deals, and delays in clinical development programs.
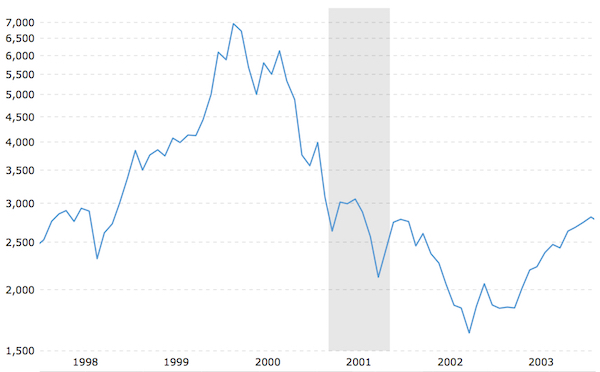
NASDAQ Market History – January 1998 to January 2004. Markets were sharply down following 3DP’s IPO in 2000, suppressing the valuation of the company. Gray bar indicates recession period of 2001.
3DP Startup
I was approached by a headhunter in 1993 to found a new structure-based design company based on research ongoing at Johns Hopkins university. The lead venture backers were HealthCare Ventures Inc., headed at the time by Wally Steinberg, who claimed fame as the inventor of the Reach™ toothbrush for Johnson & Johnson. Vertex Pharmaceuticals was in the news during this period, and I have no doubt served as motivation for HealthCare Ventures. Our key lead and BOD member from Healthcare was Hal Werner, who I always viewed as the brains behind the organization. At the time, one prevalent HCV strategy was to effectively use Series A startup funds to fund research at a university lab that would cede IP rights to a startup undertaking commercial development of the technology. Although this model had been used successfully with Human Genome Sciences, I recommended a broader approach based on my already extensive efforts in structure-based drug design at DuPont CRD, DuPont Merck Pharmaceuticals and Sterling Winthrop Pharmaceuticals Research Division.
I was able to assemble a world-class Scientific Advisory Board at 3DP that provided a wide reach over many areas in structural biology and who remained active in evaluating and helping direct he company’s research strategy. The 3DP SAB included Jeremy Berg (then at Johns Hopkins Department of Biophysics), Michael Levitt (Department of Structural Biology at Stanford and subsequently 2013 Nobel Prize laureate in Chemistry), Victor Marder (Hematologist and Professor of Medicine at UCLA), Clarence Schutt (Professor of Chemistry at Princeton), and Chairman Don Wiley (Professor of Chemistry at Harvard).
Over the lifetime of 3DP, I served as and Founder and CEO, and then President and Chief Scientific Officer of the company. My first recruit as a startup was Scott Horvitz as CFO, who joined from another HealthCare startup, Magainin Pharmaceuticals Inc. I was soon Joined by Roger Bone from Merck, who was initially to run crystallography, and Richard Soll from Wyeth Ayerst to run chemistry. A bit later I was joined by Dimitris Agrafiotis who had worked on the retrosynthesis computational platform LHASA with E.J. Corey at Harvard. Dimitris’s group built the DirectedDiversity combichem computational support platform.
Structure-Based Design and Collaborative Discovery Programs
3DPs initially identified two structure-based drug design programs as internal discovery programs. The first involved the development of orally active inhibitors of thrombin, an enzyme that plays a key role in blood coagulation. We capitalized on the advantage that thrombin produced excellent crystals that could be “soaked” with inhibitors to produce the structures of thrombin-inhibitor complexes fast enough to directly drive synthetic chemical lead optimization programs. Based on experience at DuPont, where we had worked on thrombin inhibitors based on boronic acid transition state analogs, we also knew what to avoid: eg. peptide-like structures with highly basic substituent functionality and the potential to interact with other serine protease active site residues. Informed by this reverse engineering principle, we evolved a series of inhibitors that we were able to out-license to Wyeth Ayerst for collaborative development. Ultimately, under a change in Wyeth research management, Wyeth walked away from the development program in lieu of paying a multi-million dollar milestone to 3DP. Although this situation did not make the investors happy, there was little 3DP could do. 3DP consequently took its lead anti-thrombotic compound through Phase 1 clinical studies on its own. The compound showed good safety and pharmacokinetic properties and we were able to out-license the program to Johnson and Johnson. It is key to note here that JnJ had had an internal anti-thrombotic agent discovery program. Although a competitive internal program can sometimes inhibit a collaborative discovery deal, in my experience partner conviction about the therapeutic validity of a target is the best catalyst to getting a collaboration deal.
Our second initial structure-based discovery program was focused on finding small molecule drugs that would specifically bind to Major Histocompatibility Complexes (MHCs) for inflammatory and autoimmune disease applications. Much had been learned about MHC antigen presentation through pioneering X-ray structural work carried out in Don Wiley’s lab at Harvard. However, in contrast to thrombin, routine production of the desired MHC crystals was problematic. Finally, a Nature paper by Merck investigators describing the MHC-CLIP peptide exchange mechanism made clear that a simple competitive displacement inhibitor was probably not going to work, so the program was abandoned.
While we could sometimes interest potential partners in 3DP-defined discovery targets, many technology-based deals were focused on (usually very difficult) targets defined by a partner. For example, our first combichem lead optimization deal with Merck KGaA in Darmstadt, Germany was directed towards a difficult membrane-associated target where Merck scientists had identified an initial lead. The Merck KGaA chemists wanted us to carry out a lead optimization program using combinatorial chemistry approaches and demanded that the resulting compounds lack any residual “handles” as required for synthesis on a support substrate. This requirement was clearly a matter of principle rather than necessity, but our chemists came up with inventive ways to address the issue. We were also required to set up a biological assay as a surrogate for the more complex cellular assay used in Darmstadt. We could never really get the SAR results from the 3DP and Merck-Darmstadt assays to completely agree, and Merck KGaA scientific management eventually decided not to develop the compounds that we produced during the collaboration.
Directed Diversity & The Transformative Deal That Never Happened
In 1995, 3DP was issued its first patent (of 6) for “System and Method of Automatically Generating Chemical Compounds With Desired Properties”. The patent described a process that used computational methods to control chemical synthesis of compounds with desired chemical properties, particularly using robotic chemical synthesis methods. The process described closely mirrored 3DP’s discovery process that virtualized vast chemical libraries that could be synthesized using robot-assisted, parallel chemical synthesis, and then utilized an iterative feedback optimization process to optimize potency and desied drug-like properties. At the time, this was a new concept that was very different from classical medicinal chemistry practice. Our patent lawyer, Michael Q. Lee at Sterne Kessler Goldstein & Fox, was a pioneer in the development of computer assisted method patents. The patent received a lot of attention, since it was directed at controlling a process (parallel synthesis of multiple compounds using combinatorial chemistry methodology) that had been the basis of several new biotech companies. Combichem technology, particularly when combined with newly emerging high-throughput screening technology, was under active evaluation by most major pharma companies to enhance the productivity of their discovery pipelines.
Jim Cavanaugh, an ex-SmithKlineBeckman executive and partner at Healthcare Ventures assisted us in making several meetings with SmithKline Beecham (SKB) scientists and SROne, SKB’s venture investment arm (SKB merged with Glaxo in 2000 to form Glaxo SmithKline – now GSK). Discussions were proceeding, although given the novelty of the 3DP IP position, many SKB chemists were skeptical that process control IP could be applied to medicinal chemistry. SKB’s SROne venture arm frequently made venture investments alongside research and development collaborations made between biotechnology companies and SKB, a synergy that worked in everyone’s interest. To help a potential 3DP deal along, Jim arranged a dinner with Jan Leshley (CEO of SKB), George Poste (Head of Research SKB), and Brian Metcalf (Head of Discovery Chemistry SKB). The dinner went badly since discussions had progressed without prior discussions with Poste, who consequently killed the 3DP deal. Instead Poste went on to invest in Orchid BioSciences, a company spun out of the Sarnoff Labs that had developed a system for performing parallel chemistry in micro-channeled glass plates. Since chemistry is highly unpredictable, the microchannel device never materialized as a practical parallel synthesis platform. Later the CEO, Dale Pfost would successfully transition Orchid to a completely different technology platform for genome sequencing.
Chemical Diversity and the Realities of Combichemisty
Aside from the emergence of genomic sequencing and bioinformatics methods to identify new drug targets, two additional technology advances hit the biotechnology space coincident with 3DP’s early development phase. These were high-throughput screening or HTS for the rapid evaluation of chemical compounds as biologically active “hits”, and combinatorial chemistry or combichem for the creation of large, novel chemical compound libraries. Although both technologies had many antecedents, newer advances in computer controlled robotics promised vastly enhanced productivity for both the production and screening of novel chemical compounds.
3DP responded to these challenges through the invention and development of its DirectedDiversity combichem technology and it ThermoFluor label-free HTS screening technology. Both of these technology platforms evolved through several generations at 3DP. While the development of ThermoFluor was a more or less linear process always aimed achieving higher throughput and sample sensitivity, combichem development explored a wide variety of synthetic strategies and platforms, including nanocans, resins, and microwave assisted parallel synthesis methods. Maintaining compound integrity during storage and distribution also presented issues owing the deliquescence of the solvents used to solubilize compounds for storage.
Although many research groups were advocates of strategies that screened mixed or impure samples derived from combichem synthesis (probably a hangover from the practice of screening natural products, where samples often contain mixtures of several compounds), this proved to be very disruptive to the HTS hit identification process. At 3DP our philosophy focused on avoidance of the propagation of errors, as opposed to potentially missing a minor active component in a mixture. Consequently, we built a high-throughput GLC-Mass Spec and HPLC purification platform to analyze each reaction product and, if necessary, rapidly purify compounds following synthesis. Since the location and identity of each compound was tracked by plate well from inception, we were able to rapidly triage screening hits based, in part, on the synthesis fidelity of the compound. The chemists were very satisfied with this approach, as it allowed instant identification of potential problem (e.g. relatively impure) hits once the screen was complete. Obviously, if a chemically interesting hit sample showed high potency on dose ranging studies, then there was justification for additional purification and follow-up analysis.
One recurring discussion during the evolution of the high-throughput drug discovery era centered on the limited chemical diversity of currently known drugs vs. the possible universe of known compounds with “drug-like” properties. The basic nature of this question is obvious to any medicinal chemist who readily recognizes the differences between typical synthetic molecules and natural product drugs. The current PubChem database documents chemical structures and properties of ~3x10(7) molecules. Nevertheless, the debate continues to this day regarding how much of the unexplored chemical space, comprising an estimated ~10(50) molecules, represents a genuine opportunity to discover new drugs. At 3DP, our sophisticated DirectedDiversity cheminformatics platform allowed us to generate novel synthetic libraries based on a multiplicity of desired chemical properties, either to investigate chemical diversity or to create “target-focused” libraries whose properties were biased to known compounds modulating a specific target class (e.g GPCRs, kinases, serine proteases, metalloproteases, etc.). This strategy seemed to work for virtually all of the “druggable” target classes attempted. Will completely new types of molecular architecture derived from the space of ~10(50) unexplored molecules produce drugs for currently intractable targets? Perhaps. Clearly one major current trend is to exploit the assets of repositories like PubChem to find hits for new targets that can eventually be elaborated into novel chemical leads, and eventually marketed drugs.
Genomics Driven Drug Discovery
Throughout its evolution, 3DP did a variety of collaborative discovery and chemical library out-licensing deals using either its DirectedDiversity combinatorial chemistry technology or synthetic combichem libraries. In addition to Johnson & Johnson to whom 3DP had licensed its thrombin inhibitor program, additional partners included Berlex, Heska, Aventis Crop Science, DuPont Agricultural Products, and Boehringer Ingelheim. However, 3DP’s breakthrough deal, that catalyzed its IPO in 2000, was with Bristol Meyers Squibb (BMS). As part of it’s quest for new anti-infectives, BMS molecular biologists had developed a random gene knockout strategy as a means as for defining novel antimicrobial targets. The framework of the deal, framed during discussions with David Floyd, Head of Discovery Chemistry at BMS, anticipated an extensive program finding leads for a multiplicity of targets in parallel using 3DP’s integrated technology platform. ThermoFluor, a label-free ligand screening technology that required no specific knowledge of a target protein’s enzymatic function, was well suited to this task, although we rapidly learned varying solution parameters to optimize assay parameters, as well as pre-screening with a 3000 compound “probe library” of common biochemical cofactors and substrates. In the end, we discovered that microbial targets generally produced many fewer progressable leads that targets derived from eukaryotes. We formed the impression that these proteins were effectively “Teflon coated”. In a more modern context, this behavior most likely reflects unique properties of the protein’s energy landscape evolved by prokaryotic proteins that causes them to reject binding of “foreign” small molecules.
G-protein Coupled Receptor (GPCR) Structural Biology
3DP’s interaction with a broad base of potential pharma customers sensitized us to the great interest attached to G-protein Coupled Receptors (GPCRs) both owing to their prominence among established drug targets and the identification of new orphan GPCrs emerging owing to the rapid advances in genome sequencing occurring in the 1990s. At the time, 3DP was operating at the state-of-the-art with respect to technology for protein X-ray structure determination, including synchrotron access through the IMCA consortium at Argonne national Lab and a 3DP facility for robotic setup of crystallization trials and high-throughput image analysis to monitor crystal formation. Consequently, we made a proposal to the NIST ATP program to use our resources to crystallize the beta-2 adrenergic receptor (b2AR), at that time one of the most studied GPCR receptors. We were fortunate to obtain a $2 Million award from NIST to attempt he structure determination and realigned and/or added to our resources to support the effort. One of our new hires, Cliff Robinson was assigned to take the lead in expression and refolding of the GPCR, work that seemed a logical extension of work performed for his PhD with Steve Sligar at Champaign-Urbana and a postdoc performed with Bob Sauer at MIT. Although Cliff appeared to get off to a fast start, as manifest by data presented at project meetings reporting successful protein expression and refolding, doubts began to arise around mass spec data that 3DP contracted out to characterize the fidelity of its expressed proteins. As it turned out, Cliff had completely fabricated the b2AR GPCR characterization data and substituted ovalbumin (a protein of similar molecular weight) to the crystallographers for crystallization trials. The crystallographers, in turn, had crystallized the protein and were in the process of finding derivatives to phase the structure and complete the structure determination. At that point, Cliff abruptly resigned from 3DP and took a job at the University of Delaware. This overall action had many negative repercussions, not the least among the many 3DP scientists working on the project, which we were ultimately forced to wind down. At UDel, Cliff went on to submit NIH grant proposals that reiterated many of the claims and used the same falsified data as used to deceive his 3DP colleagues, for which he was ultimately discovered and reprimanded by the NIH.
The Final Evolution
3DP went public on NASDAQ (DDDP) in a Bear Stearns-led IPO in 2000 at $15 a share, giving it an initial valuation of approximately $320 Million. At this point in time, 3DP was being led by David U’Prichard as CEO. David filled many of the top management and business positions at 3DP with former colleagues from SmithKlineBeecham, which had recently merged with Glaxo to form GlaxoSmithKline (GSK). Post IPO, the strategy of company changed significantly, with much less emphasis on collaborative drug discovery and technology licensing deals, and more emphasis on internal clinical development programs. In the latter context, 3DP had in-licensed a promising compound from GSK and was preparing a new batch of drug substance (manufactured at a European contract facility) as required to complete some bridging Phase I clinical development studies. In the event, miscommunication among the development management team resulted in the synthesis of an incorrect racemate of the active compound, which was only discovered and sorted out 12-18 months later. Lacking both robust top line revenue, and with a projected cash-burning delay reaching the clinic with its in-licensed drug program from GSK, the company appeared weak in the then existing conditions of the post-2000 market downturn, and 3DP’s valuation sank accordingly. At this point, John and Johnson, with whom 3DP had formed a long-standing relationship, stepped in and purchased the 3DP at a modest premium to the public market valuation.